Hello,
I have tried to modify the example script given under SRD to calculate vacf and mean squared displacement and I am attempting to calculate the diffusion coefficient from this. However, the values are not matching and the range is different. I also included the factors necessary for a 2D simulation (details given in examples/DIFFUSE) but this is still giving an incorrect result. I am attaching my input script as well as the output graphs. Please advice!
Sincerely,
John
in.srd.pure.nofield (1.92 KB)
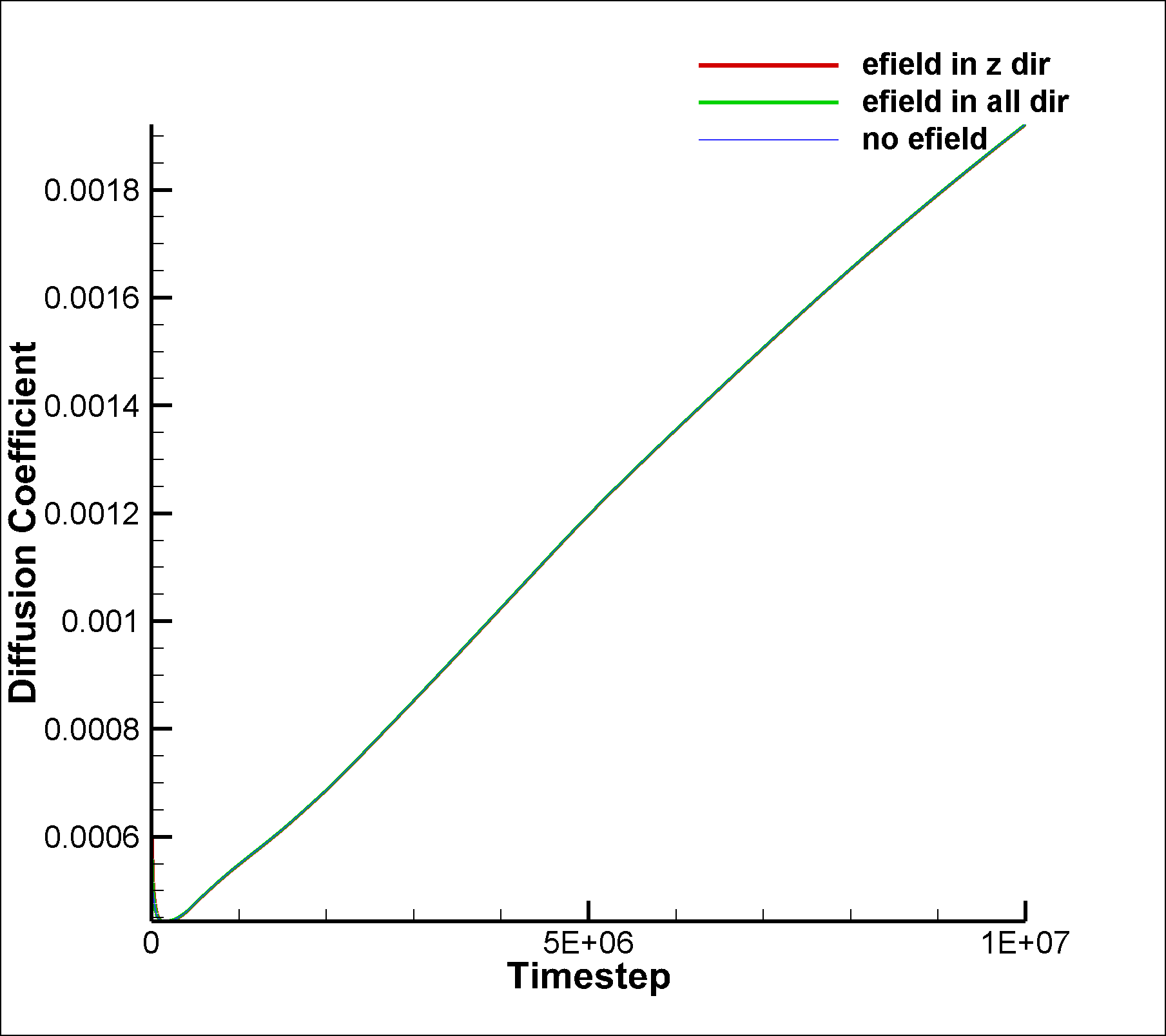
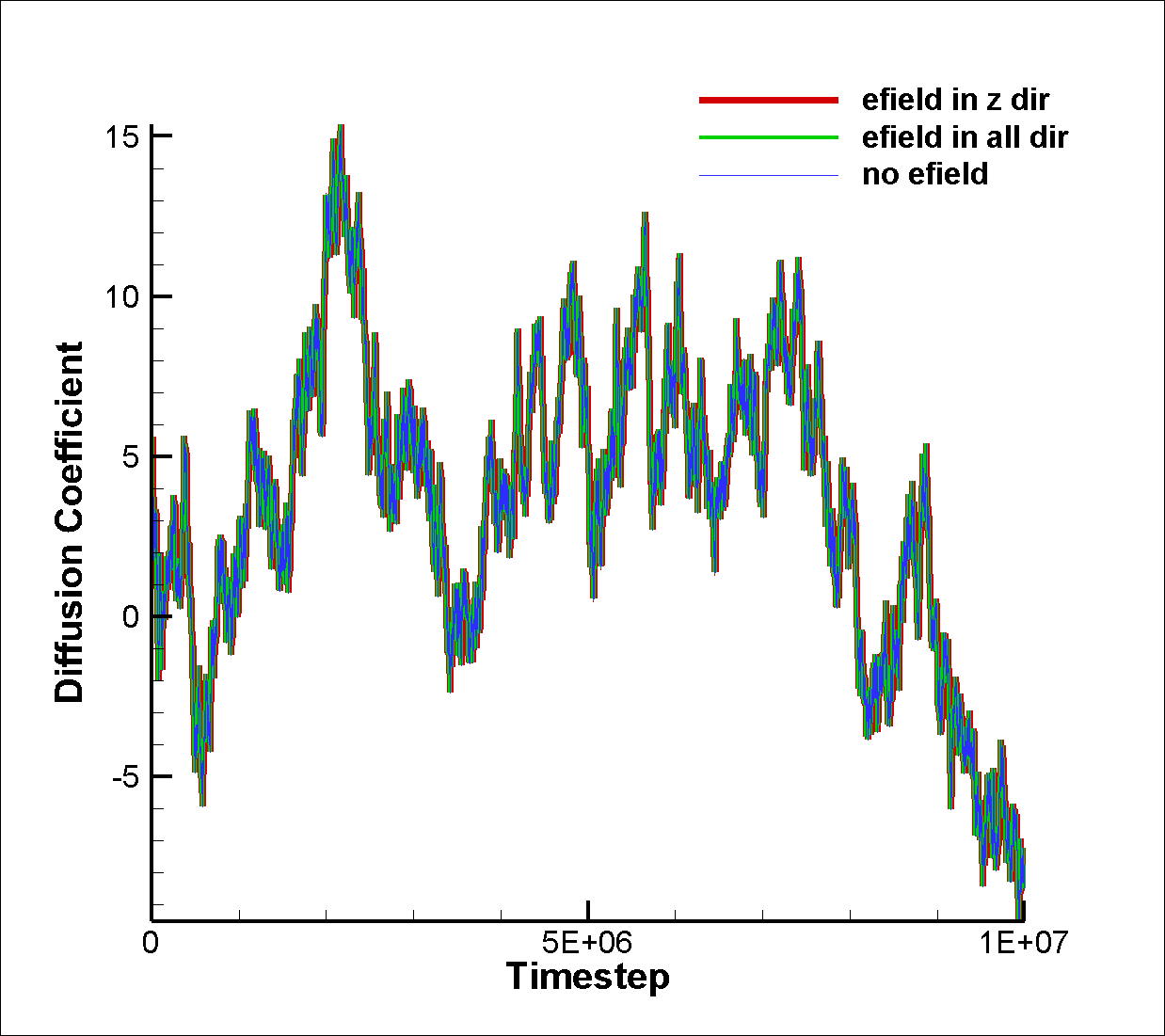
Since SRD particles do not interact with each other,
I don’t think a VACF is meaningful. Effectively
there are no velocity correlations.
When you say
MSD is incorrect, you can check yourself if
LAMMPS is producing a valid number. Just output
one or more dump files and compute the MSD yourself
at some time delta. I’m guessing you will find it
is the same. What that means for an SRD fluid
and how to interpret it, is a different Q.
Steve
I also tried the same for the mixture simulation and calculated the VACF for the non-SRD particles. In this case, there are interactions between the particles and hence velocity correlations should exist, right? However, this is not what I am observing as the diff coefficient calculated using VACF and MSD dont match.
I am once again attaching the input script and two graphs.
Sincerely,
John
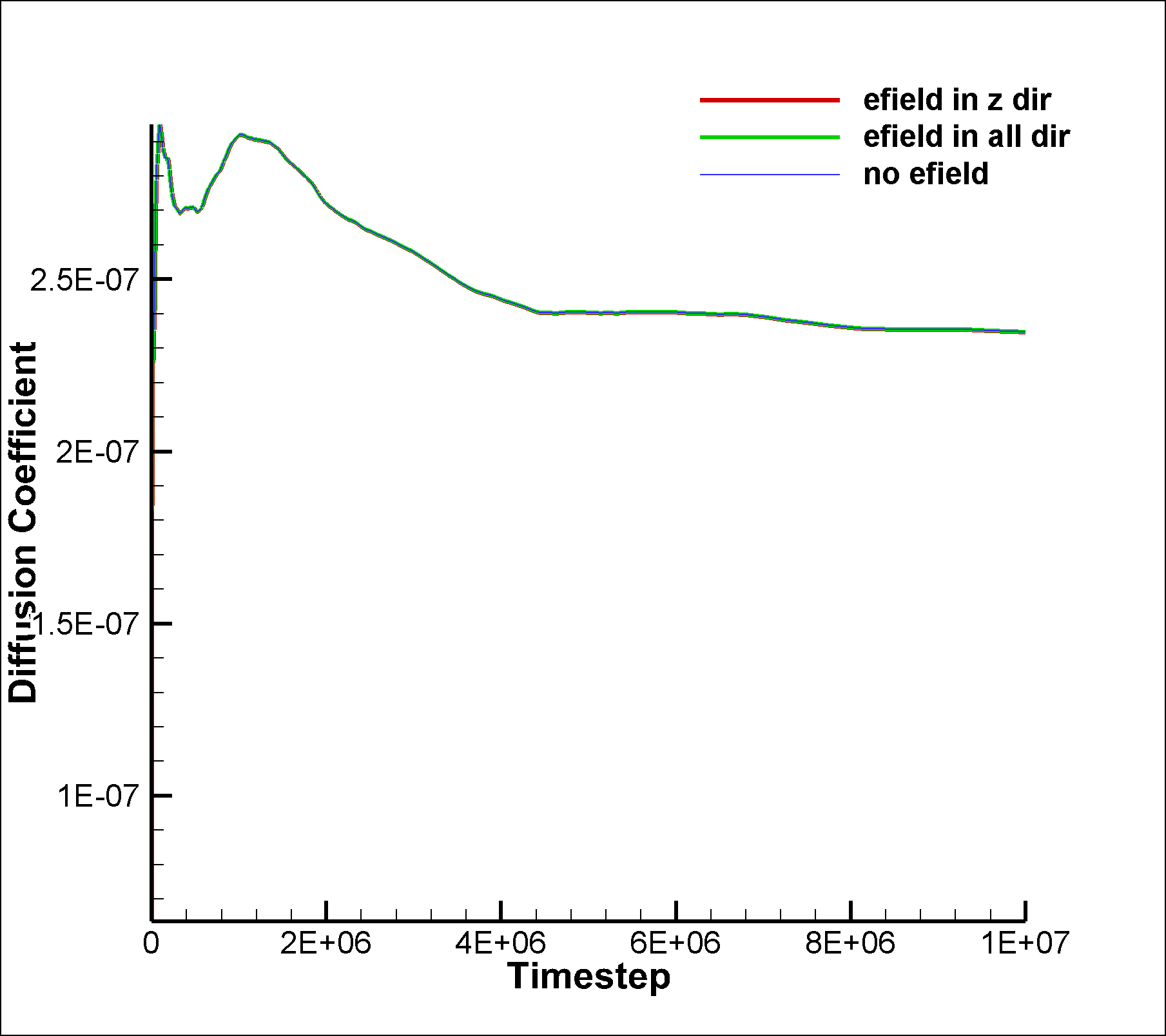
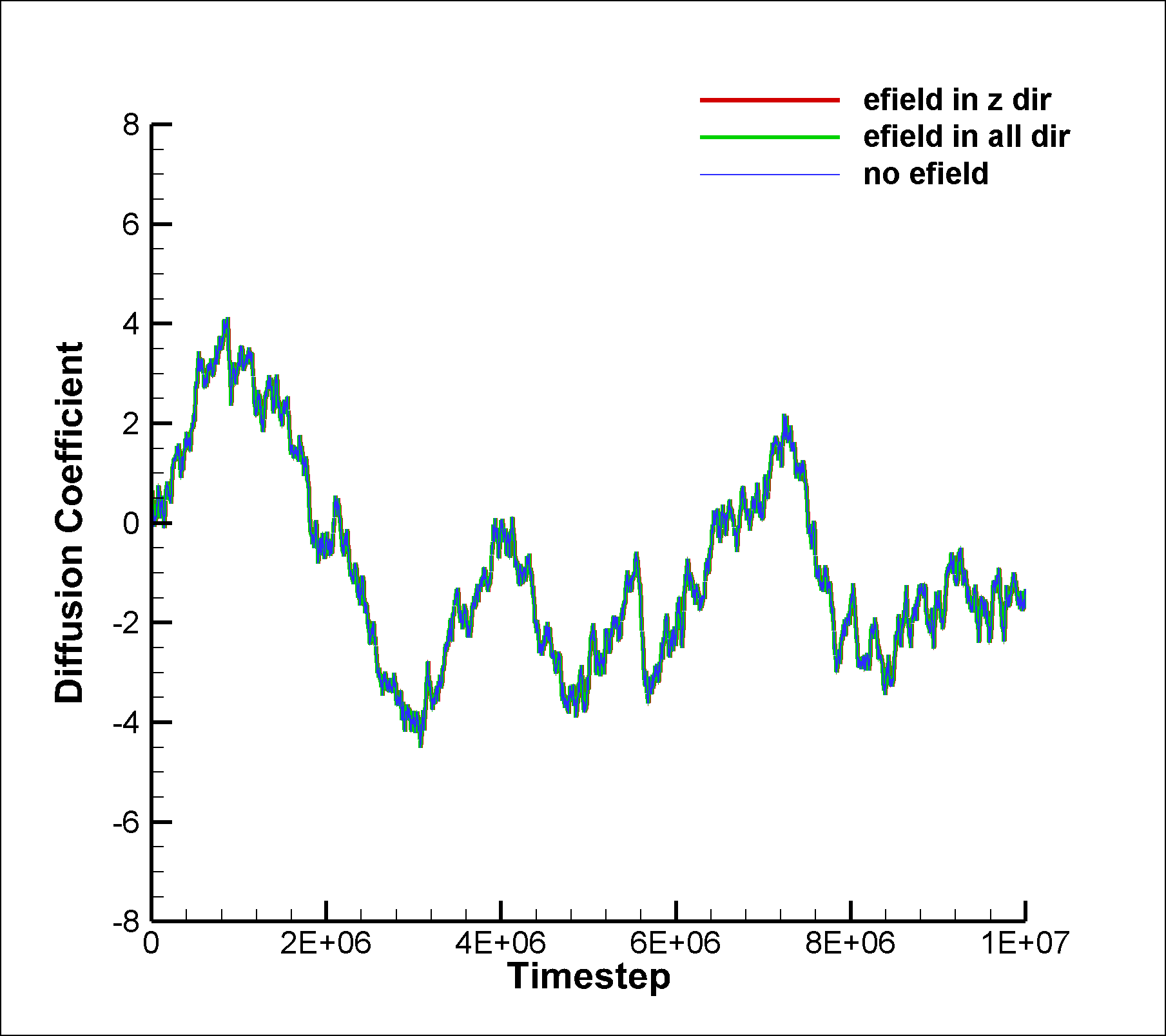
in.srd.mixture.noefield (3.12 KB)
Have you tried running the MSD and VACF input script
in examples/DIFFUSION? If so, try to see what is different
about those than your scripts. I will say that VACF is a much trickier
way to compute a diffusion coeff. You have to worry about timescales
with VACF and proper averaging and statistics, etc. MSD is
simple. There is little that can go wrong. And you can validate
with just a couple long-time snapshots, that you have a valid
MSD number as produced by LAMMPS.
Steve
Yes I ran both of them and there was a difference in the output for both methods (~0.3 using vacf and ~0.2 using msd). However, in my calculations, they don’t even have the same number of significant digits. I followed the exact same procedure as given in the examples/DIFFUSE directory and took into consideration the 2D factors as well.
What exactly do you mean by long-time snapshots?
Sincerely,
John
I mean for MSD that you are just fitting a slope to
a few data points. If the data is fairly linear,
like for a liquid, then you can get a decent slope
from 1 point (plus the origin). You can calculate
the one point from a timestep after 1000 or 10000
steps, just take the final positions minus the
original positions (e.g. from 2 dump snapshots).
So long as you pay attention to periodic boundary
crossings, you have a good estimate of the Diff coeff.
VACF has no analog to that.
Steve



