Thanks, Axel.
Sorry that it took some time before I could keep up with the issue.
I will take care not to cross-post in future, but for now, I am sending this
to both mailing lists, to share my information on lammps to vmd options.
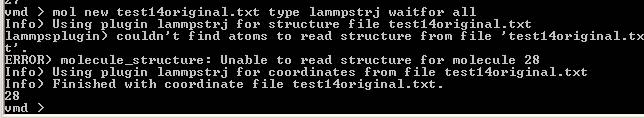
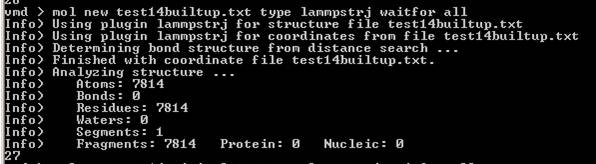
So I tried with new 1.8.7 version.
Firstly, the issue of parser bug is solved, as you said.
Secondly, I searched for topotools package so that I can test it, but no
where did I find its tutorial or option in the menu. Could you direct me to
it, please?
Thirdly, now lmp2vmd script works, but I have little issues there:
1. I don't see any visible changes happening after I implement residue tool,
in option b.
2. I can use coloring by atom name so that it gives different colors to
different atoms. This is desirable for polymer molecule, but not for polymer
and solvent molecules together if they have same atoms. So in addition, I
want to assign different colors to different molecules, even though they
carry same atoms, how to do that? that means I want to use atom name
coloring selectively for polymer chain, but for different solvent molecules
like water and methanol, I want to use different colors, even though same
atoms are also there in polymer chain. IN my data file, I have different
atom types for "such same" atoms, but belonging to different molecules, but
I don't see that vmd is taking that information in, even though in lmptoname
tool I use "H1 H2 H3 etc". I guess it assigns same color to same atom
belonging to different molecules.
So with my limited knowledge, this can be solved by, (a)either assigning
different colors to different atom "types as defined in my data file" (b)
different colors to different molecules, in addition to atom color
differentiation in polymer chain.
How to do that??
Thanks very much
Regards
Chetan