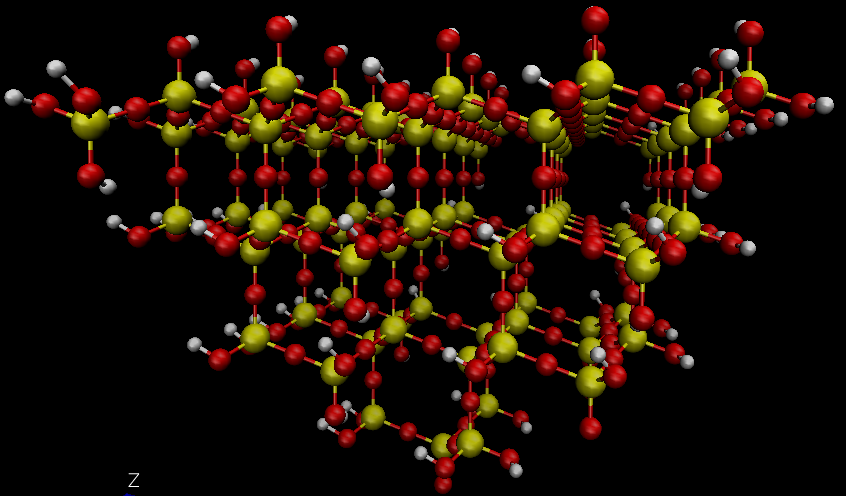
Yes, it is not required to use different atom types, but it is good practice. It can be extremely helpful to identify the different kinds of Si atoms this way during the simulation and for analysis.
Since you are building your own data files, you are free to set the charge and atom type as you will. How to construct this, should be straightforward, as when looking at the connectivity.
(See below)
, if you also use different atom types for the different kinds of O atoms.
Axel.
Dear Jewett and Axel
Thank you very much for your responds.
But my main point is how to distinguish the surface Si (with OH group) atom from the bulk (without OH group).
The easiest way is to write a short program that will loop through the list of bonds and print the ID numbers of the two atoms, whenever they both satisfy the atom-types (O and H).
To find the list of bonds you can write a for loop to read the “Bonds” section of the DATA file, or use pizza.py to read the entire data file.
Then write another loop that will use the information above to generate a list of charges (or “set atom ID charge Q” statements)
https://LAMMPS.sandia.gov/doc/set.html
If you want to write something fancier and more general, and if you are using python, you can try using “networkx” to search for subgraphs in the bond network.
https://networkx.github.io/documentation/stable/reference/algorithms/isomorphism.vf2.html
In your case, the subgraph has two bonds and one edge (O-H ). The VF2 algorithm will find a list of atom-IDs (vertices) which for all of the O-H “subgraphs” in your bond network.
If the VF2 algorithm is too slow, you can try using the following code, which is almost the same. It is written in the same style, and is much faster than VF2 for small subgraphs (like the OH group):
https://github.com/jewettaij/moltemplate/blob/master/moltemplate/nbody_graph_search.py
Andrew