Hi,
I am a PhD Student at Trinity College Dublin. I am creating this post to discuss a pymatgen wrapper for ASE-GUI that I’ve been using for a while to set up my resting state calculations for electrocatalysis in VASP.
The analysis of the surface coverage is an essential first step for a correct study of a periodic system, and until a couple of months ago, I used to calculate this by manually adsorbing H, O and OH on the surfaces, definitely missing possible adsorption sites for larger and less standard surfaces. Then, I developed a python script that calls the pymatgen.analysis.adsorption module to automatically find the X and Y coordinates of every possible adsorption site using a Delaunay triangulation with the surface atoms. This is then passed to ASE, which iteratively approaches the adsorbates from an initial distance until contact with the surface atoms, removes the structures with colliding or non-surface adsorbates and performs post-processing (constraining, adsorbing on the other side of a slab, etc). This yields a list of Atom objects containing all the possible surfaces with adsorbed species, ready for calculation. I repeat this process to obtain the resting state.
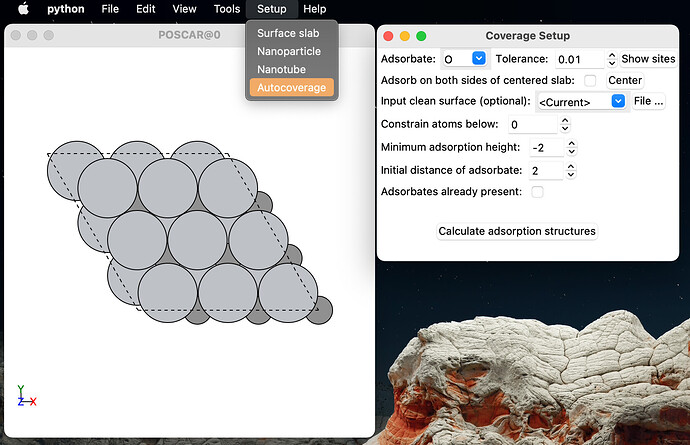
I think this is important because there is no tool available that automatically detects adsorption sites of non-standard, multi-element slabs and generates all the possible structures with any desired aqueous adsorbate. However, relying on a script made it hard to implement for the rest of my research group. That’s why I modified ASE-GUI to include this functionality:
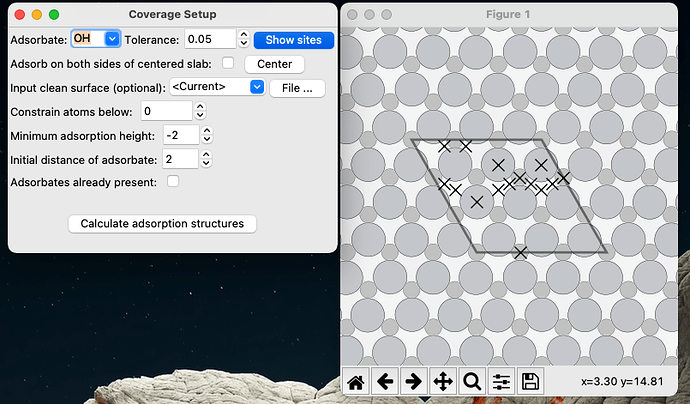
Which then allows you to see the adsorption sites schematically
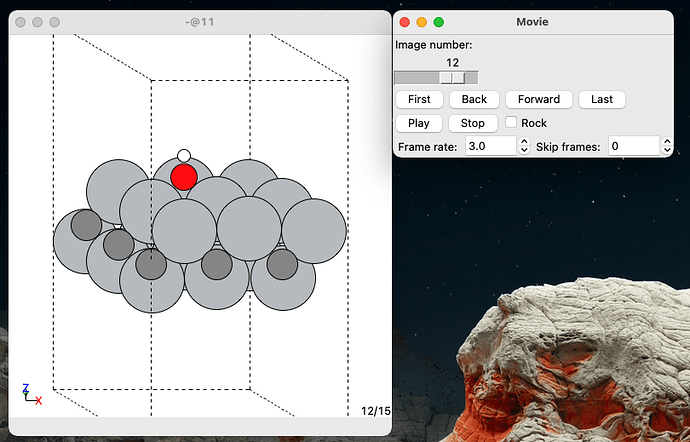
And generates a movie containing the structures for all the adsorbed species
I was wondering whether it would be feasible to merge this into the official version of ASE? It heavily relies on the pymatgen library, though, so I’d understand if not, but it would help the work in my group considerably.
Thank you very much to the devs for all the hard work, ASE has been the structure for most of my projects!
Eric