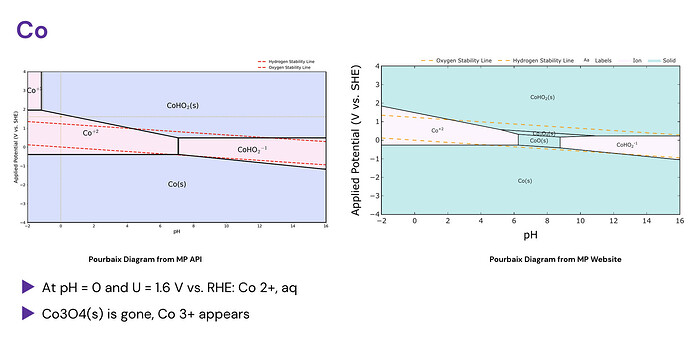
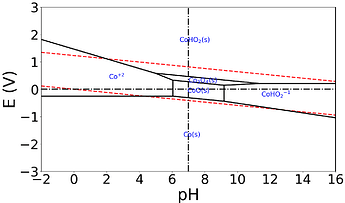
Good day, I have tried to use the MP API to generate my own Pourbaix diagrams locally, and I seem to notice from large differences between the API and the website.
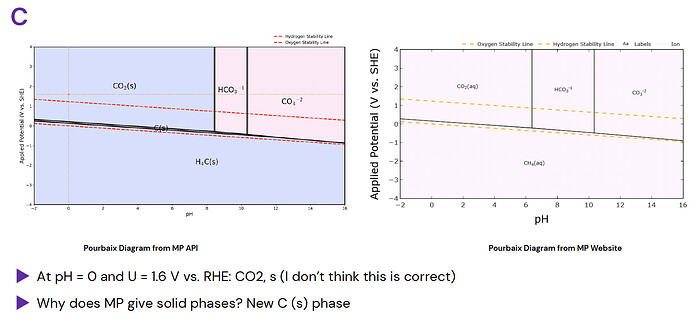
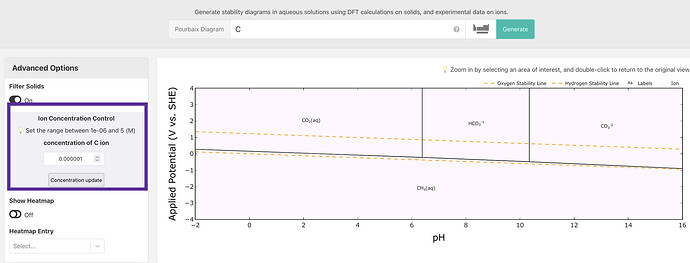
There are some diagrams with clearly mistaken phases, such as CO2 from Carbon’s Pourbaix Diagram:
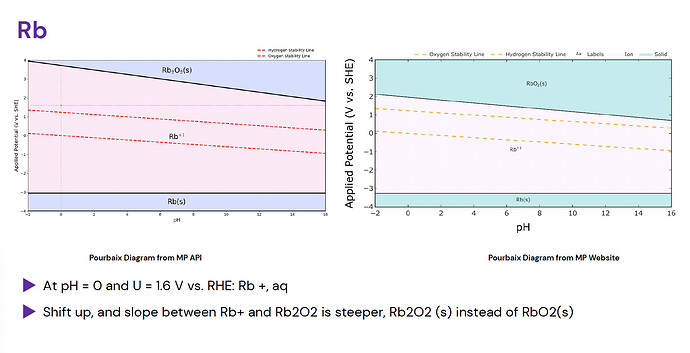
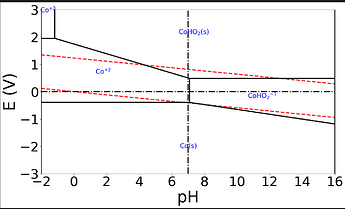
There are some patterned errors, particularly in Group 1A or 2A elements, where a different oxide is shown:
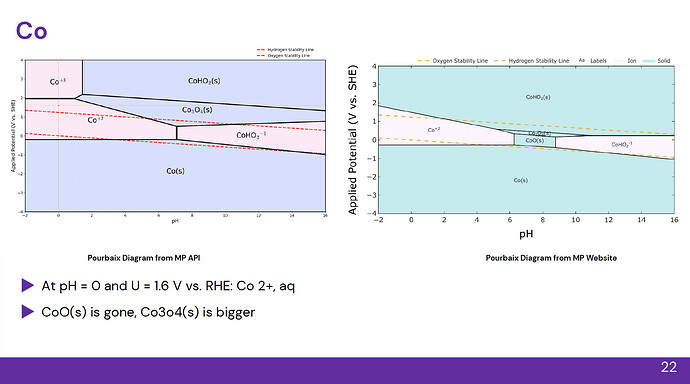
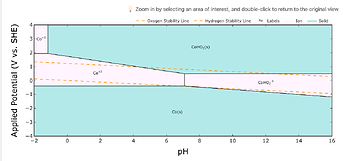
In other cases, phase sizes are vastly different:
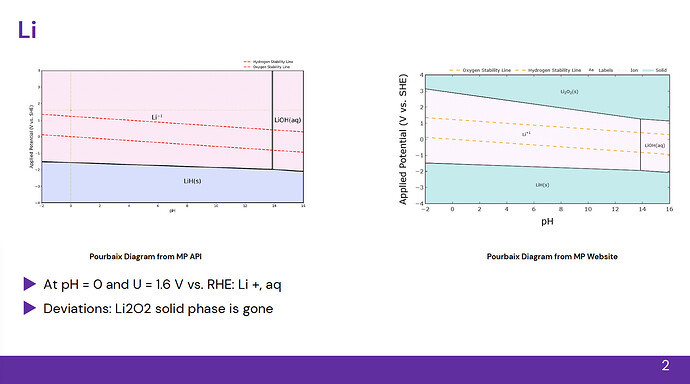
Other times, new phases appear or disappear:
Please see the code below (conducted for all elements except S, Cr, Se, Ag - for a different discussion)
from mp_api.client import MPRester
import matplotlib.pyplot as plt
import numpy as np
from collections.abc import Iterable
from matplotlib import rcParams
from matplotlib import pyplot as plt
from pymatgen.analysis.pourbaix_diagram import PourbaixDiagram, PourbaixPlotter, PourbaixEntry, MultiEntry
from matplotlib.patches import Polygon
from scipy.spatial import ConvexHulldef _is_constant(vals, tol=1e-9):
True if all values equal to a constant within tolerance
return np.nanmax(vals) - np.nanmin(vals) < tol
def remove_default_guides(ax, ph_target=7.0, v_target=0.0, tol=1e-6):
“”“Remove the vertical pH=ph_target and horizontal V=v_target lines
that PourbaixPlotter draws by default.”“”
to_remove =
for ln in ax.get_lines():
x = np.array(ln.get_xdata(), dtype=float)
y = np.array(ln.get_ydata(), dtype=float)# Identify a vertical line at pH ≈ ph_target if _is_constant(x) and np.isfinite(x).all(): if np.isclose(x[0], ph_target, atol=tol): to_remove.append(ln) continue # Identify a horizontal line at V ≈ v_target if _is_constant(y) and np.isfinite(y).all(): if np.isclose(y[0], v_target, atol=tol): to_remove.append(ln) continue # Remove the matched lines for ln in to_remove: ln.remove()-------------------------------
GLOBAL FONT: SERIF
-------------------------------
rcParams[“font.family”] = “sans-serif”
rcParams[“mathtext.fontset”] = “cm” # computer modern math look-------------------------------
Get MPR API Key and Select Element
-------------------------------
#el_list = [“Li”, “Be”, “B”, “C”, “N”, “F”, “Na”, “Mg”, “Al”, “Si”, “P”, “Si”, “Cl”, “K”, “Ca”, “Sc”, “Ti”, “V”, “Cr”, “Mn”, “Fe”, “Co”, “Ni”, “Cu”, “Zn”, “Ga”, “Ge”, “As”, “Se”, “Br”]
el_list = [“C”]for element in el_list:
with MPRester() as mpr: # 'Fe' is the element of interest; O and H are added automatically pbx_entries = mpr.get_pourbaix_entries(element, use_gibbs=300) print(f"Number of Pourbaix entries: {len(pbx_entries)}") # ------------------------------- # CREATE DIAGRAM & BASE PLOT # ------------------------------- pbx = PourbaixDiagram(pbx_entries) plotter = PourbaixPlotter(pbx) #fig, ax = plt.subplots(figsize=(10, 6)) ax = plotter.get_pourbaix_plot( limits=((-2, 16), (-4, 4)), ) # Now retrieve the figure that the plotter actually created fig = ax.figure # ------------------------------- # Auto font scaling based on size # ------------------------------- base_width = 12 scale = fig.get_figwidth() / base_width # ------------------------------------------------------ # STEP 0 — Reference lines: pH=0 (vertical) and E=1.6 V vs RHE (horizontal) # ------------------------------------------------------ # Dark yellow (choose a readable, print-safe tone) dark_yellow = "#b8860b" # DarkGoldenRod # A fine dashed pattern (short dashes) fine_dash = (0, (2, 3)) # on=2pt, off=3pt # Create the two guide lines (no legend entries) ph0_line = ax.axvline( 0.0, color=dark_yellow, linestyle=fine_dash, linewidth=1.2, zorder=4, label="_nolegend_" ) E16_line = ax.axhline( 1.6, color=dark_yellow, linestyle=fine_dash, linewidth=1.2, zorder=4, label="_nolegend_" ) # Intersection marker at (pH=0, E=1.6 V vs RHE) # Use a small solid circle; we'll make it responsive too. pt_line = ax.plot( [0.0], [1.6], "o", color=dark_yellow, markerfacecolor=dark_yellow, markersize=4.0, zorder=5 )[0] # --- Register originals so the resize callback can scale them --- # (Do this once, right after creating the artists.) if not hasattr(ax, "_resp_lines_extra"): ax._resp_lines_extra = [] if not hasattr(ax, "_orig_linewidths_extra"): ax._orig_linewidths_extra = [] ax._resp_lines_extra.extend([ph0_line, E16_line]) ax._orig_linewidths_extra.extend([ph0_line.get_linewidth(), E16_line.get_linewidth()]) # Store the point (single Line2D) and its original marker size ax._resp_point_extra = pt_line ax._orig_markersize_point = pt_line.get_markersize() #remove default lines remove_default_guides(ax, ph_target=7.0, v_target=0.0) # ------------------------------------------------------ # STEP 1 — Sample (pH, V) space and label regions # ------------------------------------------------------ pH_vals = np.linspace(-2, 16, 200) V_vals = np.linspace(-4, 4, 200) sample_points = [] sample_labels = [] for pH in pH_vals: for V in V_vals: entry = pbx.get_stable_entry(pH, V) if entry is not None: sample_points.append([pH, V]) sample_labels.append(entry.name) sample_points = np.array(sample_points) # ------------------------------------------------------ # STEP 2 — Group points by phase # ------------------------------------------------------ unique_phases = sorted(set(sample_labels)) phase_points = {ph: [] for ph in unique_phases} for (pt, ph) in zip(sample_points, sample_labels): phase_points[ph].append(pt) # ------------------------------------------------------ # STEP 3 — For each phase, take convex hull to form polygon # ------------------------------------------------------ for ph, pts in phase_points.items(): pts = np.array(pts) if len(pts) < 3: continue # need ≥3 points to form hull hull = ConvexHull(pts) poly = pts[hull.vertices] # polygon vertices # Color by solid vs aqueous/ion if "(s)" in ph: facecolor = (0.55, 0.65, 1.0, 0.35) # blue else: facecolor = (1.0, 0.75, 0.90, 0.35) # pink patch = Polygon(poly, closed=True, facecolor=facecolor, edgecolor="black", linewidth=0.8, zorder=0.3) ax.add_patch(patch) # ------------------------------------------------------ # STEP 4 — Double the label sizes (keep black) # ------------------------------------------------------ for txt in ax.texts: txt.set_fontsize(txt.get_fontsize() * 1.25) txt.set_color("black") # ------------------------------------------------------ # STEP 5 — Fix legend using the default red dashed water lines # ------------------------------------------------------ dashed = [ln for ln in ax.get_lines() if ln.get_linestyle() == "--"] if len(dashed) >= 2: dashed[0].set_label("Hydrogen Stability Line") dashed[1].set_label("Oxygen Stability Line") #ax.legend(frameon=False, fontsize=14, loc="upper center", ncol=2) ax.legend(frameon=False, loc="upper center", ncol=2) # Create legend leg = ax.legend( frameon=False, fontsize=14, loc="upper left", bbox_to_anchor=(0.76, 1.10) ) # Store baseline anchor so we can scale relative to it ax._legend_baseline_anchor = (0.76, 1.10) # ------------------------------------------------------ # Axes labels and ticks # ------------------------------------------------------ ax.set_xlabel("pH", fontsize=18) ax.set_ylabel("Applied Potential (V vs. SHE)", fontsize=18) ax.tick_params(labelsize=14) plt.tight_layout() # ------------------------------------------ # Dynamic font + line scaling on window resize # ------------------------------------------ base_width = fig.get_figwidth() # initial width in inches # Store original line widths once if not hasattr(ax, "_orig_linewidths"): ax._orig_linewidths = [ln.get_linewidth() for ln in ax.get_lines()] def on_resize(event): new_width = fig.get_figwidth() scale = new_width / base_width # ---- Text scaling ---- ax.xaxis.label.set_fontsize(14 * scale) ax.yaxis.label.set_fontsize(14 * scale) for t in ax.get_xticklabels() + ax.get_yticklabels(): t.set_fontsize(14 * scale) leg = ax.get_legend() if leg: # Scale legend text for txt in leg.get_texts(): txt.set_fontsize(12 * scale) # Scale legend line thickness legend_lines = leg.get_lines() # Store original legend line widths once if not hasattr(leg, "_orig_linewidths"): leg._orig_linewidths = [ln.get_linewidth() for ln in legend_lines] for ln, orig_w in zip(legend_lines, leg._orig_linewidths): ln.set_linewidth(orig_w * scale) for txt in ax.texts: # phase labels txt.set_fontsize(14 * 1.25 * scale) # ---- Line scaling (NEW) ---- for ln, orig_w in zip(ax.get_lines(), ax._orig_linewidths): ln.set_linewidth(orig_w * scale) # ---- NEW: scale the reference guide lines & intersection marker ---- if hasattr(ax, "_resp_lines_extra") and hasattr(ax, "_orig_linewidths_extra"): for ln, w0 in zip(ax._resp_lines_extra, ax._orig_linewidths_extra): ln.set_linewidth(w0 * scale) if hasattr(ax, "_resp_point_extra") and hasattr(ax, "_orig_markersize_point"): ax._resp_point_extra.set_markersize(ax._orig_markersize_point * scale) fig.canvas.draw_idle() fig.canvas.mpl_connect("resize_event", on_resize) # ------------------------------------------------------ plt.savefig('Pourbaix_' + element + '.png')
I am also attaching a Google Drive link which is a comprehensive comparison between MP API and MP Pourbaix Diagram’s Webpage. This PDF covers all elements except for f-block elements and those without Pourbaix Diagrams on the Webpage.
Thank you, and I am looking forward to your response.