My ovito version is 3.10.6.
This is the version of ovito library:
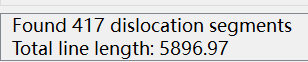
In the GUI, OVITO tells me that the model has a total of 417 dislocations and that the total length of these dislocations is 5896.97:
In the python script, I use the following statement to count how many dislocation lines are in the current model file and their total length.
from ovito.io import import_file
from ovito.modifiers import DislocationAnalysisModifier as DXA
pipeline = import_file("dump_20000.shear", input_format="lammps/dump",
sort_particles=True)
num_frame = pipeline.source.num_frames
pipeline.modifiers.append(DXA(input_crystal_structure=DXA.Lattice.FCC))
data = pipeline.compute(frame=0)
result = 0.0
for line in data.dislocations.lines:
result += line.length
print(len(data.dislocations.lines), result)
After running the above code, I get the following output:
![]()
This means that when I call the DXA algorithm from the OVITO library via python, it tells me that there are 426 dislocation lines in the current model, and that the total length of these dislocation lines is 5856.63.
And there is a slight discrepancy between such a result and the one I get in the OVITO GUI, so I am confused, is there something wrong with my python code? Or is this just normal?
Finally, as I’m a new user so unfortunately I can’t upload my dump files, but I’ve found after a few tests that these differences can be triggered consistently in the version of OVITO I provided earlier.