Dear all LAMMPS Users,
I am writing to share an issue that I have encountered while configuring a bilayer graphene system in LAMMPS, and to request your insights and suggestions.
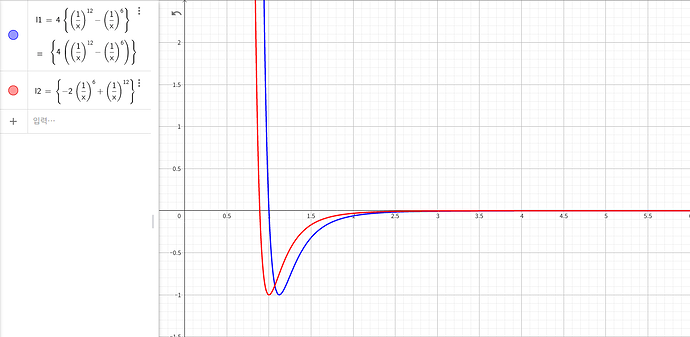
As you may know, bilayer graphene has a distance of 3.34Å between two layers. However, when I configured my system and minimized it in LAMMPS, I found that the distance between the two layers was about 4.57Å. I suspect that this discrepancy may be due to an error in the parameters that I am using, but I have not been able to identify the source of the problem. I uploaded my reference below this post. I modified the parameter to use proper value. Because in LAMMPS, pair_style lj/cut command used 12-6 LJ potential, but the paper used 6-12 type.
I am hoping that some of you may have encountered similar issues or have insights into what could be causing this discrepancy. If you have any suggestions or recommendations on how I can resolve this issue, I would be very grateful.
Thank you in advance for your help and insights.
LJ potential para.pdf (1.3 MB)
This is my parameter modified version, type 1 is carbon atom, type 2 is Si atom.
pair_coeff 1 1 0.105 4.622601348
pair_coeff 1 2 0.205450724 4.571787923 16.0012577305
pair_coeff 2 2 0.402 4.820974497
bond_coeff 1 469 1.4
angle_coeff 1 63 120
dihedral_coeff 1 0 7.25 0 0
improper_coeff 1 5 180
Additionally, this is minimization part of my LAMMPS inputfile,
---------- Initialization ----------
units real
atom_style full
pair_style lj/cut 14
boundary p p p
bond_style harmonic
angle_style harmonic
dihedral_style opls
improper_style harmonic
special_bonds lj 0.0 0.0 0.5
---------- System definition ----------
read_data /home/jungwan/lj_project/bilayer/Si_bilayergraphene.data
include /home/jungwan/lj_project/parameters/PARM_Si_graphene.lammps
region supported_graphene block INF INF INF INF -0.5 0.5
region sample_graphene block INF INF INF INF 2.85 3.85
group substrate region supported_graphene
group sample region sample_graphene
group Si type 2
---------- System Relax ----------
fix freeze substrate setforce 0.0 0.0 0.0
dump 1 all atom 1 minimized.lammpstrj
minimize 1.0e-10 1.0e-10 200000 200000
write_data minimized_coordinated.data pair ij