Hi,
Hi,



Hi,
I am using fix rigid to glue beads in a rod. I relax my system to a zero
force configuration using 0-temp Langevin dynamics. when I checked a pair
of rigid rods with 3 beads in each rod, I found that the fix constraint is
not exactly kept, and the error is in the sixth digit. the rods are
slightly bending.this doesn't make sense. if fix rigid is working correctly (and i have
seen many many cases where it does), then this cannot happen because of how
it is implemented. the only explanation i can think of is that your data
file is not correctly set up, i.e. it doesn't assign molecule ids properly.
does the output actually confirm that you have 4 four rigid bodies?
I put four parallel rods (short or long) in a box which are free floating
in space. then I relax them and calculate sum of the bead_wise Virial_x. My
hypothesis is that since the bundle is not connected to anywhere, the
virial in x direction has to be zero. These are the result I get from
LAMMPS: a = spacing between beads, sigma = box units, box length = 60this is the LAMMPS input file:
atom_style molecular
read_data data_4r3B.dat
mass 1 1
mass 2 1fix fixRods all rigid molecule
fix fixLangevinOSC all langevin 0.0 0.0 10.0 1234 zero
yes
fix fixNVEOSC all nve
timestep 0.05
compute PotentialEOSC all pe
fix fixPEOSC all ave/time 1 1 1 c_PotentialEOSC file
pot_relaxing_eps_0.txtcompute peratomOSC all stress/atom pair
compute pOSC all reduce sum c_peratomOSC[1]
c_peratomOSC[2] c_peratomOSC[3] c_peratomOSC[4] c_peratomOSC[5]
c_peratomOSC[6]thermo 10
thermo_style custom step temp pe etotal press xlo xhi
thermo_modify flush yes
dump dumpCus all custom 10000000 4rods3B.xyz.* type x
y z fx fy fz c_peratomOSC[1] c_peratomOSC[2] c_peratomOSC[3] vx vy vz
dump_modify dumpCus format "%d %20.15g %20.15g %20.15g %20.15g %20.15g
%20.15g %20.15g %20.15g %20.15g %20.15g %20.15g %20.15g" sort idpair_style lj/cut 1.58
pair_coeff 1 1 0.001 1.0 1.1225
pair_coeff 2 2 0.10 1.0 1.58
pair_coeff 1 2 0.001 1.0 1.1225
neigh_modify exclude molecule all
log log_manualRelax_eps_0.dat
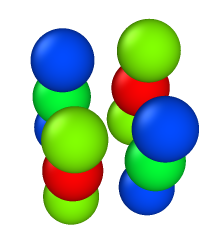
run 100000001) 4rods, 3beads in each, a/sigma = 1.0 —> LAMMPS Virial_x = -2.9e-13 -
This case LAMMPS giving zero value for virial.
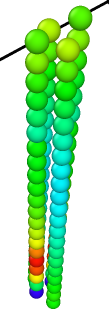
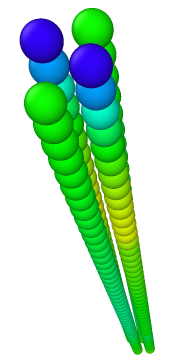
beads are colored based on Virial_x
3) 4rods, 21beads in each, a/sigma = 1.0 —> LAMMPS Virial_x = -5.16e-5 -
LAMMPS reports non-zero virial in x direction.
4) 4rods, 51beads in each, a/sigma = 1.0 —> LAMMPS Virial_x = -0.00153210
- LAMMPS reports non-zero virial in x direction.
5) 4rods, 51beads in each, a/sigma = 0.4 —> LAMMPS Virial_x =
-0.0289228943Looks like error depends on BOTH Number of beads in a rod, and a/sigma.
In all case studies, all the beads are type 2 particles.
Do you have any insight on why I would get non-zero virial for a free
floating bundle? Any help would be appreciatednobody can say anything unless you provide a simple to use test case that
completes quickly. right now, there are too many additional unknowns (what
version of LAMMPS, what OS/compiler, what hardware) that could all have an
impact.
axel.


