Hi everyone,
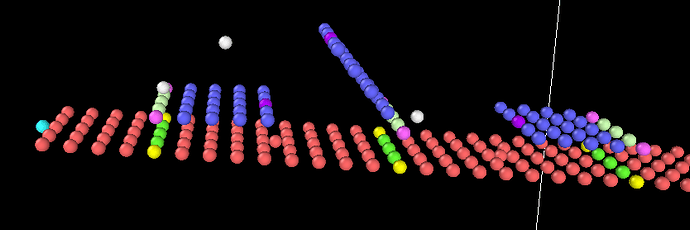
I’ve been trying to model a system which starts with a bond between the right brown bead and the right blue bead. Then,
- when the left brown bead gets closer to the blue one, a bond forms between the left brown and the left blue bead with a probability of 30%
-when there is a bond between the left blue bead and the left brown bead, then we break the bond between the right brown bead and the right blue bead.
To do this, as suggested by Will Pisani, I am considering a bond/react fix. However, I’m getting the following error:
ERROR on proc 3: Bond/react: Excessive iteration of superimpose algorithm (src/USER-MISC/fix_bond_react.cpp:1219)
Any suggestions of what could be going on? Attached I’m sending my setup.
Thanks!
Sebastian
in.lammps (3.14 KB)
data.system (9.68 KB)
reaction0.dat (13.1 KB)
rxn1_stp1_map (1.5 KB)
reaction1.dat (13.1 KB)
I am noticing that your reaction templates contain all the atoms in your system. Reaction templates should only contain atoms participating in the reaction (bond/react will search the system for reaction sites). So, in your case, looks like your templates should have only three atoms each.

Hi Jacob,
Thanks for your reply. I tried to do that change but I still get the same error.
The original atom numbers are 43, 48, 150 and 175.

reaction0.dat (330 Bytes)
reaction1.dat (329 Bytes)
rxn1_stp1_map (84 Bytes)
Hi Sebastian,
Now, the issue is that all atoms must be connected to a ‘bonding atom’ (which we are now calling ‘initiator atoms’). You could model the reaction in two steps (probably the easiest way). Or, you could add a ‘dummy bond’ between those two atoms that stay put.
Jake

Hi Jacob,
Thanks for your reply. I set up the simulation again and now I don’t run into that issue anymore. However, I’ve been running lately into:
"Atom affected by reaction too close to template edge"
I found online a suggestion for this
"This means an atom which changes type during the reaction is too close to an 'edge' atom defined in the superimpose file. This could cause incorrect
assignment of bonds, angle, etc. Generally, this means you must include more atoms in your templates, such that there are at least two atoms
between each atom involved in the reaction and an edge atom."
But didn't understand it very well nor how to adapt it to my model. What I'm currently doing is kind of like playing "tag":
- the atom on the left is tag (Type 10); when it gets close to type 8, it switches with the atom on the right (type 7).
- in the reaction, I also change type 8 to type 11 because if not the reaction becomes stuck on a loop here
Any suggestions about how to fix this error are more than welcome. Thanks and merry Christmas!


So, are you still having issues, or did my last suggestion solve the problem? Note that the quickest way to solve the problem is to include the relevant files.
Hi Jacob,
I’m still thinking of alternatives of solving the problem, thanks for asking!
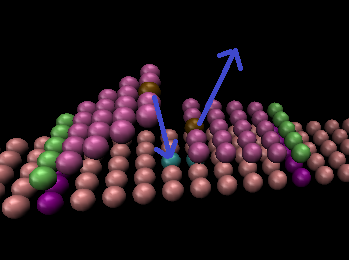
Please find attached my files. Everything starts with a freely diffusing left slab and two slabs bound to the surface, middle and right.
The idea is that, when the left slab reaches the middle slab, the left slab will bond to the surface and the middle one will become free, all this with a probability p (for now I set this to 1).
Now, the middle one will diffuse. If it reaches the right one, the same thing will happen with a probability of p. But if it reaches the left one, it can only free the left one with a small probability of p/10, for example.
To do this, I wanted to change the type of the left sensor and have two states: probability p (if type 8), and probability p/10 (if type 11)
It’s a reversible reaction with kf>kb (right now with two steps, but I want to generalize it to N) and I’m currently struggling with the fact that we need things to be bonded in order for fix bond/react to work, and the slab in the middle will then have to be bonded to the sensor in the left and the sensor on the right.
Thanks,
Sebastian
reaction0.dat (268 Bytes)
in.origami (3.39 KB)
data.system (11 KB)
reaction1.dat (268 Bytes)
dna_tether_equil.lammpstrj (11.4 KB)
rxn1_stp1_map (101 Bytes)
Hi Jacob,
I got this to work, the solution was to include all atoms in such a way that the edge atom does not change type, as suggested.
Thank you very much!
Sebastian