Dear all,
I have met several problems.
Problem 1: I use the lammps to simulate the adatom on the surface,
when I set “boundary p p fs”, and add “fix addbox addbox
wall/reflect zhi”, however there is still a mistake about “ERROR: Lost
atoms: original 3096 current 3092”. How can I handle the problem?
there are multiple reasons for "losing" atoms.
you could have unphysical starting conditions
(high potential energy) or use too large a time
step. in both cases, single atoms may be accelerated
too much for the neighbor list update. the best
way to check this, is to write out coordinates very
often and visualize the resulting trajectory.
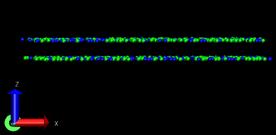
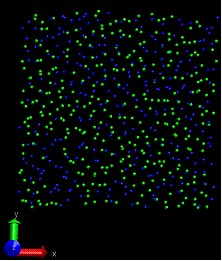
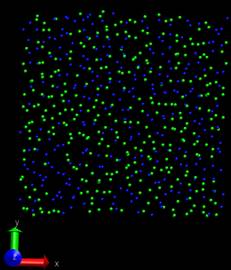
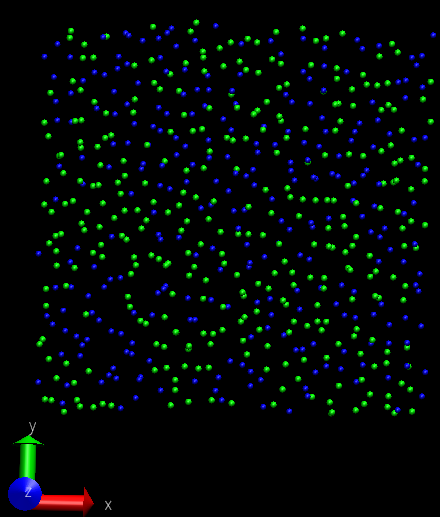
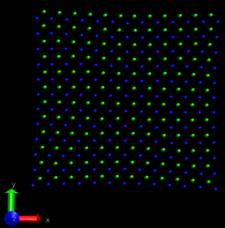
Problem2: I want the adatom atoms first in arrayed configuration, and
they will become random during the simulation, however, they just
contain the arrayed configuration and move downward to the surface
(surface is partially fixed and boundary is p p p). When I set the
surface free and p p p boundary, then I can obtain what I want―arrayed
adatom atoms become random and deposit to the surface, only the adatom
near the top are always there. If I change the boundary to p p fs,
then, everything is as wrong as the situation in (surface is
partially fixed and boundary is p p p). I just confused about that.
Could anyone give me some suggestions?
i am confused as well. what are "arrayed" atoms?
perhaps you can create some simple graph or example
input illustrating your problem.
Problem3: Maybe it is the problem of the computer cluster, I have
found that in dump file, some frame will contain more atoms than the
normal number. I guess this is correlated with mpi or something
associated with the parrellal calculation. Have anyone met the same
problem?
this does not make any sense at all. there cannot be more atoms
unless you create them during the simulation. are you sure you
are not confusing outputs from different calculations?
cheers,
axel.