Thanks a lot for your response.
First I tried with pdb file to take atom's coordinates and time I didn't
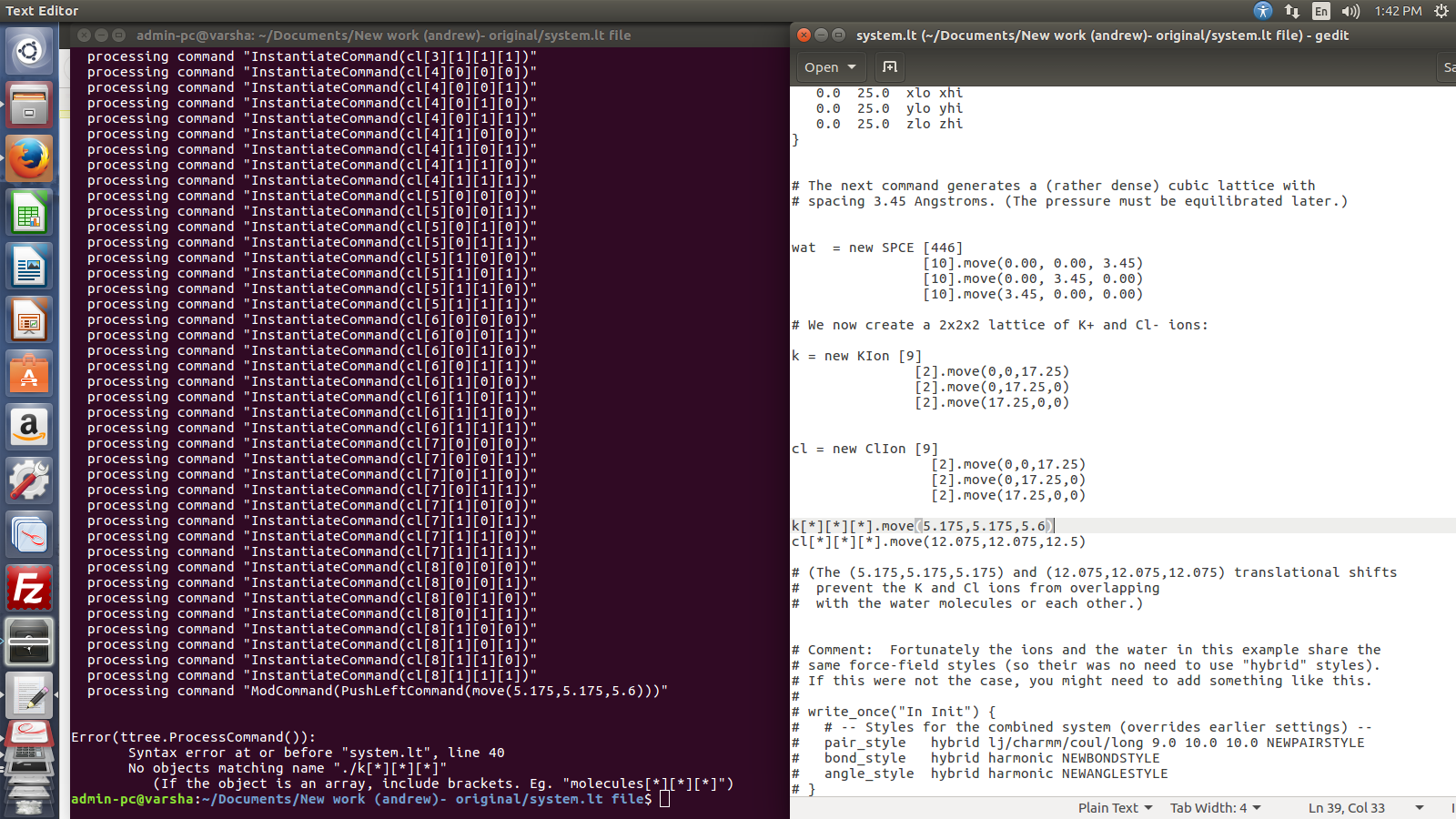
mention "move" command in the script (system.lt)
But when I received the output file (system.data), I found some errors in
atom's coordinates. The reason of this is - that pdb file was incorrect
which I used.
So I tried it with "move" command so that I can get atom's coordinates in
the moltemplate itself.
I have not any choice if I will remove "move" command from system.lt .
I have not added a way for moltemplate to modify the coordinates of
atoms that you are reading from a PDB (or XYZ or RAW) file.
I suggest you should either fix the problem with your PDB file, or use
.move() commands for the entire system.lt file.
Alternatively, you CAN use moltemplate to generate a system where SOME
of the coordinates come from a PDB file, and other coordinates are
created using moltemplate's "move()" commands, although it's a little
more tricky.
Below I sketch how you could do that. (I have not actually tried the
procedure that I outline below, but it should definitely work.
However you may have to correct mistakes in my awk commands. The awk
commands should fit on one line, but they may get split into 2 lines
in the process of emailing them.)
1) Create an LT file (eg, "system_excerpt.lt") containing only the
molecules whose coordinates are missing from your PDB file. (This LT
file will contain the molecules whose position you control with
"move()" commands.) Use moltemplate.sh to generate a DATA file for
these atoms:
moltemplate.sh system_except.lt
2) Then use a text editor to cut out the "Atoms" section of the data
file (eg "system_excerpt.data"), and use "awk", or "cut"+"paste" or
"pizza.py", to extract the columns corresponding to the atom
coordinates from that data file, and finally save them in a new simple
3-column text file (eg "coords_excerpt.raw").
A faster way to accomplish the same thing is to use the following commands:
extract_lammps_data.py Atoms < system_excerpt.data > Atoms.txt
awk '{print $4" "$5" "$6}' < Atoms.txt > coords_excerpt.raw
3) Do the same thing with your PDB file to create a 3-column text file
with the PDB atom coordinates (eg "coords_pdb.raw"). One way to do
that is using awk:
awk '/^ATOM |^HETATM/{print substr($0,31,8)" "substr($0,39,8)"
"substr($0,47,8)}' \
< YOUR_FILE.pdb > coords_pdb.raw
4) THEN append the text from your 3-column coordinate file extracted
from your DATA file to the 3-column coordinate file extracted from
your PDB file, and save the result in a new file (eg "coords.raw".
cat coords_excerpt.raw coords_pdb.raw > coords.raw
5) Finally use your "system.lt" file (the file which includes all of
the molecules in your system including some which came from your PDB
file), listed in the order that they appear in the ".raw" file you
just created (hopefully). Run moltemplate on this file:
moltemplate.sh -raw coords.raw system.lt
This should generate files you can use with LAMMPS (eg"system.data",
"system.in.init", "system.in.settings") However I'm not sure this
helps with your issue.
Cheers
Andrew