Hi all,
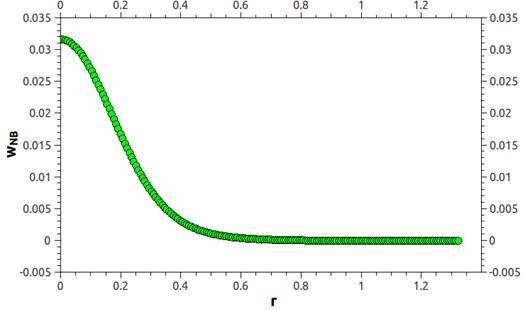
i wish somebody can help on that one. I’m trying to run DPD simulations of coarse-grained polymer chains. I’m using pair_style table and bond_style table commands in order to define the conservative force and pair_style dpd/tstat to thermostat my system. The tabulated values for bonded and non-bonded interactions were taken from the potential of mean force obtained after performing MD simulation at the coarsed-grained level under study. In order to combine pair_style table and dpd/tstat, i’m using pair_style hybrid/overlay. Following the tips given by LAMMPS documentation, i started by using linear interpolation on pair_style table, i matched the N of the pair_style equal to the tabulated N, and i tried to use pair_write command in order to produce a plot of what the final interpolated potential looks like but without succeeding (i get an error saying “Pair style does not support pair_write”. Is there any way to solve this issue?). Since DPD uses soft potentials, beads can overlap, so my potentials inner cutoff is very close to zero value (if i use an inner cutoff of zero, LAMMPS though me an error saying “Invalid pair table cutoff”). So, after running my simulation, everything seems to be fine, but after several thousand steps i get and error saying “Pair distance < table inner cutoff”. Coincidence or not, I realized that this error appears just before the simulation reaches the target temperature. What i realize about this error is that particles moved to close, to a distance lesser than what the table can afford. Below i show how the non-bonded potential in LJ units i’m using looks like (it was taken from Guerrault X, Rousseau B, Farago J, J. Chem. Phys. 121, 13, 6538-6546 ), and also parts of my table file:
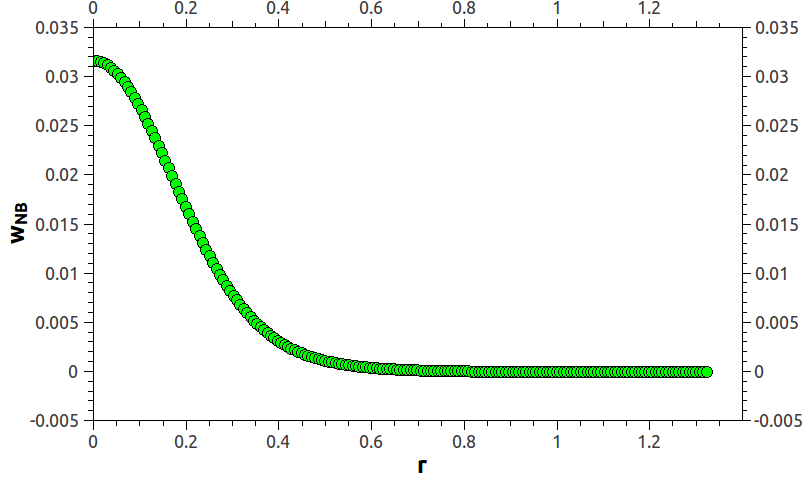
PMF_PE_NB
N 181 RSQ 0.00001 1.32159
1 0.00001 0.03166 0.00000
2 0.00734 0.03163 -0.07680
3 0.01468 0.03155 -0.15316
…
179 1.30690 0.00000 0.00000
180 1.31424 0.00000 0.00000
181 1.32159 0.00000 0.00000
I wonder if anybody can give me any tip or advise on how i can solve this “Pair distance < table inner cutoff” issue.
Thanks so much for the time.
Many regards
Sacha