Please always report which LAMMPS version you are using!
I don’t see any problems with the mixing of pair coefficients.
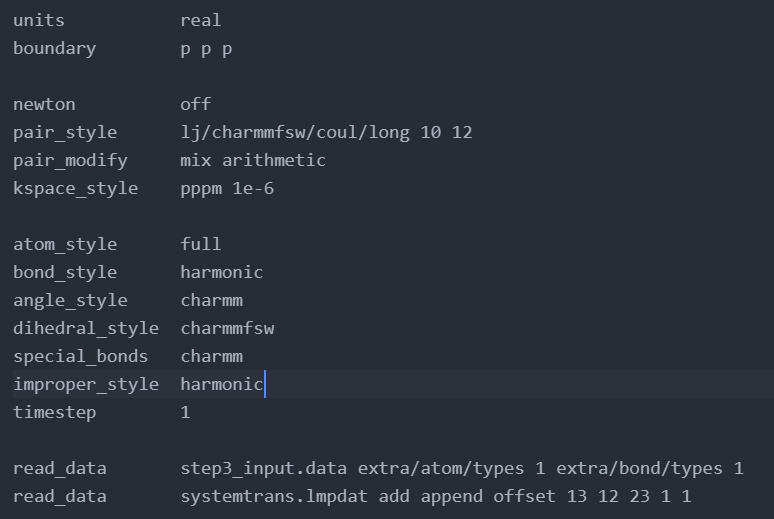
If I add a run 0 to your quoted input I get with the latest LAMMPS version (3 Aug 2022):
Generated 13 of 91 mixed pair_coeff terms from arithmetic mixing rule
ERROR: All bond coeffs are not set (src/bond.cpp:85)
So the problem is not the mixing of the pair coefficients, but that you needlessly reserve an extra bond type in the first read_data command, but never provide any bond coefficient data for it. This is confirmed by looking at the info_coeffs.txt file. After removing that unused bond type, the run 0 command can proceed and compute the systems energy.
However, there are other, quite serious problems:
WARNING: System is not charge neutral, net charge = 195.16 (src/kspace.cpp:327)
This is very bad. Systems with long-range coulomb should be neutral or at best within less than one charge unit away from it to reduce errors. LAMMPS will continue to run, but the validity of the results is questionable. You should check on the charge assignments or include a suitable number of counter ions (and solvent?) to make your system neutral.
WARNING: Bond/angle/dihedral extent > half of periodic box length (src/domain.cpp:940)
That suggests that there may be something incorrect with your topology data.
Similarly, your potential energy for the non-bonded interactions and bonds is extremely high:
Step Temp E_pair E_mol TotEng Press
0 0 1.1019126e+11 3528348.7 1.1019478e+11 4.3144838e+11
That suggest that there is something not quite right with the geometry, possibly with the box information. You should probably check out each data file independently before combining them.
To avoid issues with the read_data command expanding your box unexpectedly, I also strongly suggest to make certain that both data files have the same box sizes.