Hi there,
Until now, I have been producing povray renders of atoms objects using the ASE GUI Render Scene tool.
I am now trying to automate this process and make it easier to standardise views.
I am using the method described on File input and output, particularly in the save_pov.py and saving_graphics.py example files:
write('atoms.pov', atoms, **generic_projection_settings,
povray_settings=povray_settings).render()
With generic_projection_settings and povray_settings have been defined as:
generic_projection_settings = {
'rotation': '90x,80y,90z',
'radii': 1.0,
'colors': None,
'show_unit_cell': 1,
}
povray_settings = {
'transparent': False,
'camera_type': 'perspective',
'textures': ['jmol' for atom in atoms],
'canvas_width': 500,
'camera_dist': 100,
}
There doesn’t seem to be much guidance on this in the ASE documentation or in the POVRAY documentation, so I think I have a few questions on this method.
-
Settings in generic_projection_settings and povray_settings don’t appear to be interchangeable between dictionaries or combinable into one dictionary. What is the difference between the two and is there a list of possible settings/keywords which each one accepts? I have found lots of possible settings, but none which differentiate between the two.
-
My IDE also doesn’t recognise the .render() method and there doesn’t seem to be anything said about this anywhere. I think that this produces the .png files from the .pov and .ini files, but can anyone explain what the .render() method is actually doing/using?
The main thing I am trying to do with this function is to visualise trajectory files. I currently render the atoms object in each .traj frame in POVRAY and then combine the png files into a gif using the convert linux command.
-
On the off-chance there is one, is there a quick solution to visualising trajectory files in this way? e.g. in the ASE GUI, there was the “render all frames” option. Is there a way to “render all frames” with this method without calling the function individually for each frame in the trajectory file as I have been doing?
-
Related to question 2, when I call it individually for each frame, the resolution changes for each image. I’m assuming this is because I have set the canvas_width setting, but when I don’t set that option, it produces 64x38 pixel images… Is there a way to make the image sizes consistent?
![]()
- Another problem I had was that the atoms objects sometimes went off one side of the images. My solution to this was to try to zoom out using the camera_dist setting. However, this doesn’t seem to change anything in the final png images. 50 seems to be a default value in these examples, but even when I significantly change it, the only change seems to be the camera location in the .pov file, whilst the resulting .png images are identical, perfectly fitting the atoms object in the first frame. The only time I have got the image to change was using the ‘perspective’ camera type and putting the camera obscenely close to the atoms to get a very warped image. Can anyone explain what the camera_dist setting does and suggest how I can zoom out in my images in the same way I was able to in the ASE GUI?
This is the sort of image I am getting out of this method:
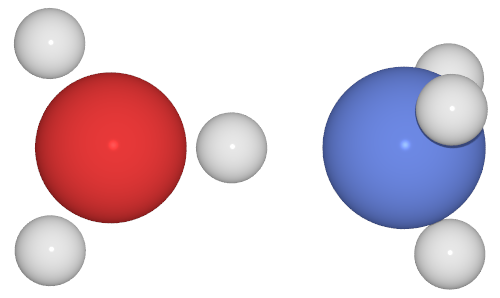
The only change has been setting camera_dist to something like a value of 1, which gives this:
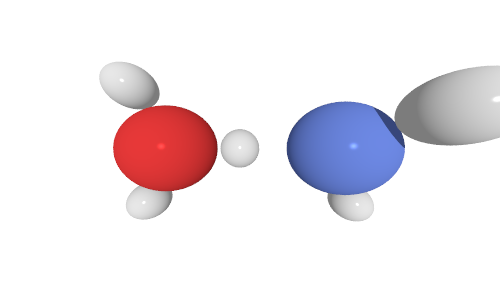
Alternatively, if anyone knows of a helpful resource with lots of help with this, that would be amazing. I have searched for a while for one, but only found the examples in the ASE documentation (which, while helpful for showing the code, don’t explain too much…) and the POVRAY documentation (which certainly has a lot of info, but is hard to sift through and, in my mind, doesn’t match up well with what I can see ASE doing…) A simple walkthrough of the methods and possible settings would be fantastic!
I think that’s probably enough questions for now, although I’ll probably have more later.
I’m really grateful for any help with this.
Thanks so much,
Matt