Dear forum members,
I have encountered a problem while using the ConstructSurfaceModifier function in my code. Specifically, I am trying to construct a surface using certain particles from the file “dump_initial_config.colloid”(see below) using the alpha shape method. The IDs of these particles are listed in the file “special_cluster_atoms”. Although these particles do not form a compact shape, they do form a cluster.
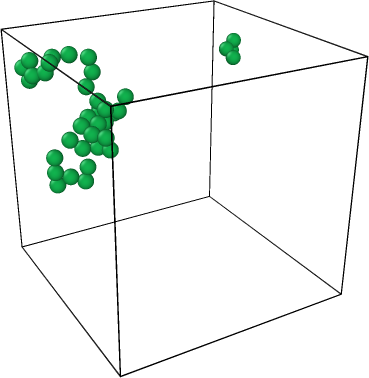
To construct the surface using these particles, I have used the option “only_selected=True” in the ConstructSurfaceModifier function. Additionally, I want to check if other particles are inside the surface by using the option “map_particles_to_regions=True” in the same function.
Following the example in the OVITO documentation, I attempt to count the number of particles inside the surface. However, I encounter the following error:
in_surface = np.count_nonzero(surface_region[‘Filled’][data.particles[“Region”]])
IndexError: index 5 is out of bounds for axis 0 with size 5
Below is the relevant portion of my code:
import ovito
import numpy as np
def main():
special_atoms_id = np.loadtxt(“./special_cluster_atoms”, skiprows=9)
dump_file = “./dump_initial_config.colloid”
pipeline = ovito.io.import_file(dump_file)
pipeline.modifiers.append(ovito.modifiers.ExpressionSelectionModifier(expression="%s"convert_ovito_command(special_atoms_id)))
pipeline.modifiers.append(
ovito.modifiers.ConstructSurfaceModifier(method=ovito.modifiers.ConstructSurfaceModifier.Method.AlphaShape,
radius=1.6,
identify_regions=True,
map_particles_to_regions=True,
only_selected=True))
data = pipeline.compute(0)
surface_region = data.surfaces['surface'].regions
in_surface = np.count_nonzero(surface_region['Filled'][data.particles["Region"]])
def convert_ovito_command(atoms_id_list):
head = ‘ParticleIdentifier ==’
joint = ‘||’
command = ‘’
for atom_id in atoms_id_list:
command = command + head + str(atom_id) + joint
command = command[:-2]
return command
if __name__ == ‘__main__’:
main()
Interestingly, when I increase the radius in the alpha shape method (e.g., changing it from 1.6 to 3), the error disappears. Could someone kindly provide possible solutions to this issue?
By the way, if I have a list of particle IDs that I want to select, I currently use the convert_ovito_command function to generate an expression. Is there a more convenient method to achieve this?
Thank you in advance for your assistance.
dump_initial_config.colloid (159.8 KB)
special_cluster_atoms (385 Bytes)