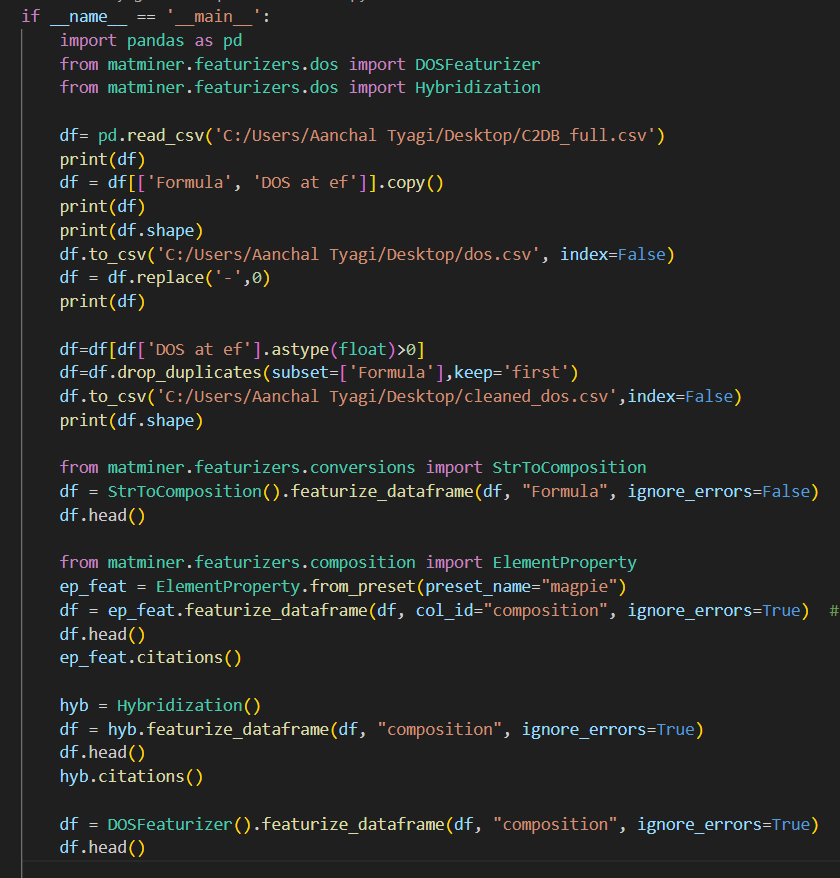
I am facing issues as I could not figure out how to use DOSFeaturizer or any other structure based featurizer for the datasets from C2DB database.
I have problem in getting the structures of the compounds from different database.
There are many examples where composition based featurizers were used…but no example for structure bsed featurizer is avavilable.
Please help me out.
@ardunn
Hi @photon01 thanks for posting here!
So to use structure featurizers or DOS featurizers, you need to put the data into a form matminer can understand. Matminer uses pymatgen under the hood for representing structures and density of states objects, so if your data can be transformed to pymatgen objects, you can use the matminer featurizers.
Essentially, your workflow will be this:
Raw data → Pymatgen → Matminer featurizers
If you are doing this en masse, matminer has utilities that can do this without the intermediate pymatgen step (for some cases). So your workflow would be:
Raw data → Matminer conversion classes → Matminer featurizers
Pymatgen - Structure data
You can browse some of the pymatgen I/O here: https://github.com/materialsproject/pymatgen/tree/master/pymatgen/io
There are interfaces for a lot of input/output files from computational software.
Similarly, you can do simple stuff like reading CIF files just from the structure class using Structure.from_file. Looking at the data on the C2DB briefly, it looks like they use ASE (i.e., from the Band Alignment example in their docs), which can be converted to pymatgen structures (see ASEAtomsAdapter here).
Matminer - Structure data
If you want to do this en masse, especially from a dataframe, you can use the ASEAtomsToStructure conversion featurizer. Then use any of the structure featurizers that are applicable.
Bandstructure
As far as bandstructure data goes, you may be on your own a bit as I’m not familiar with C2DB. Seems like they have some Db entries for stuff like the CBM/VBM but not the entire bandstructure or DOS as any kind of programmatic object?
It may be worth contacting them to see if they have programmatic access to the full BS/DOS, particularly in Pymatgen format. If you can convert the DOS to pymatgen format, you can use the matminer featurizers.
Also, if you want some more examples, check out here: https://github.com/hackingmaterials/matminer_exampleshttps://github.com/hackingmaterials/matminer_examples
Though some of these examples are a bit old!
@ardunn
thank you so much.
I will try and see what I get.
@ardunn Sir
I took DOS without SOC column from C2DB databse and featurize it using the code:
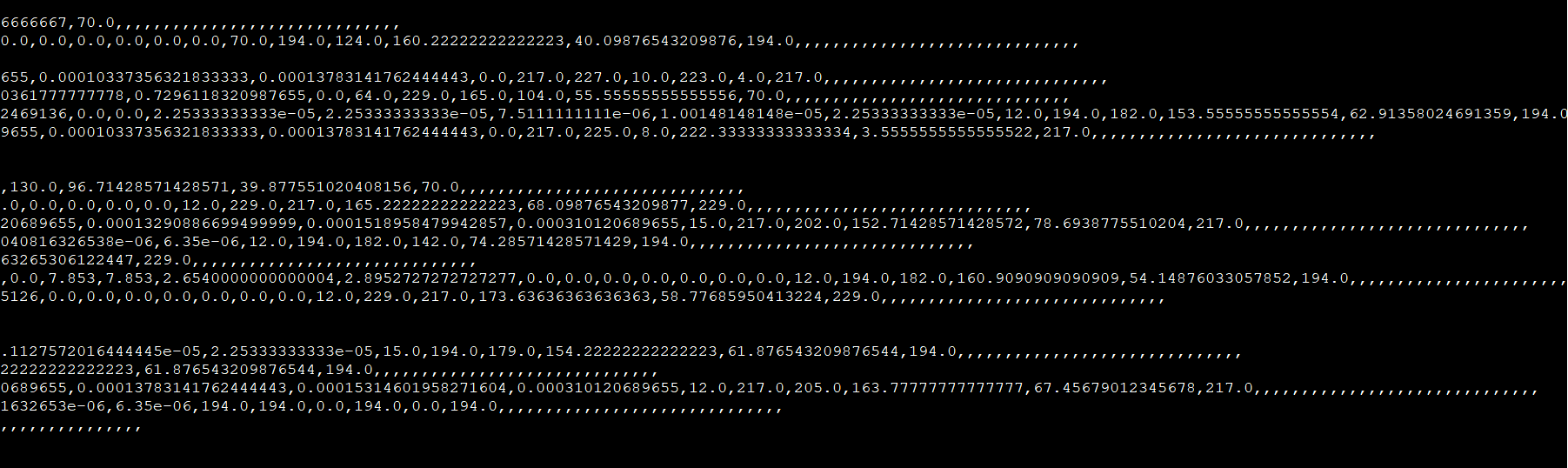
and the result I got is:
Last 30 feature columns are empty. The names of last few columns are
cbm_sf,cbm_pd,cbm_pf,cbm_df,vbm_s,vbm_p,vbm_d,vbm_f,vbm_sp,vbm_sd,vbm_sf,vbm_pd,vbm_pf,vbm_df,cbm_hybridization,cbm_character_1,cbm_specie_1,cbm_location_1,cbm_score_1,vbm_hybridization,vbm_character_1,vbm_specie_1,vbm_location_1,vbm_score_1
How can this problem be resolved?
Hi @photon01 I think this has to do with the format of the DOS objects. You need to have the full density of states as a pymatgen DOS object (pymatgen.electronic_structure.dos module — pymatgen 2023.5.10 documentation) or Bandastructure object (pymatgen.electronic_structure.bandstructure module — pymatgen 2023.5.10 documentation) to use the BS/DOS featurizers in matminer. Are you able to convert your data to these formats?
For reference, what I think is happening here is:
- Your objects in the dataframe are not proper pymatgen objects for featurization with matminer
- The ignore_errors=True is set for your featurizers
- The featurizers throw errors because the BS/DOS objects are not correct, matminer catches and ignores these errors, and your output dataframe is a bunch of empty entries.
I’d also recommend turning OFF ignore errors (ignore_errors=False) and seeing what happens. If it throws errors, please paste the stack trace in this thread so we can debug together.
Sure Sir.
I will share the problems and errors soon.
@ardunn
Hi @photon01 @ardunn
Were you able to convert the ASE dos from C2DB database to Pymatgen dos object which Matminer can understand?
The documentation says it needs Complete Dos and Structure. I was able to get pymatgen structures but DOSFeaturizer and Hybridization are still not working.
I am facing a similar problem converting dos data (from c2db) to CompleteDos. Just wondering if anyone found a solution yet?
Thanks in advance.
@ehlee
I needed data for projected dos which is actually missing in database so I simply went for electronic structure calculations using VASP and it wasn’t time-consuming at all. I chose VASP because Pymatgen CompleteDos can easily be obtained from vasprun.xml while documentation for ASR is still missing.