Hi all,
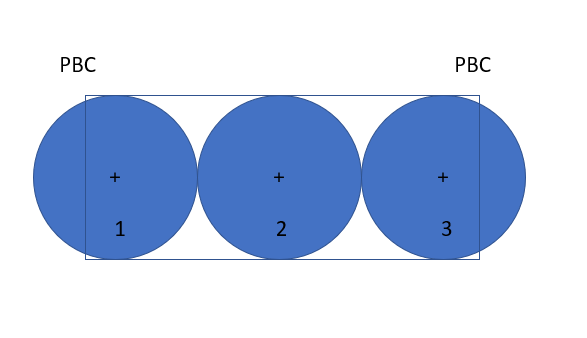
I wrote a custom granular pair potential a few months back that works well for a wall/particle bounded system but when tested on a granular specimen with periodic walls on all sides, I get a segmentation fault at the first timestep due to one of the atom vector variables not extracting the correct quantity. Upon closer inspection, I noticed that failure happens to granules at the boundary.
For example, the pairwise interaction of granule 9153 (as obtained from tag[i]) and 9599 (as obtained from tag[j]) fails because these two granules are at a periodic boundary. In the input file, I have a periodic boundary at z=0 and z=0.8m. Here are the x,y,z coordinates of granule 9153 and 9599 in the input data file:
9153: 4.0941922423730681e-02, 1.9419058391353688e-02, 3.2053353230316953e-07
9599: 3.8314403595514357e-02, 1.5487448856834229e-02 8.0007039234515032e-01.
When I extract these granular coordinates during the pair potential evaluation, my coordinates are roughly the same but are accounted for periodicity (note the z dimension of granule 9599):
9153: 4.0941925114364e-02, 1.9419072048767e-02, 3.11622e-7
9599: 3.8314403595514e-02. 1.5487448856834e-02, -8.0690400520e-5.
Now, specifically, my pair potential has a dynamic radius quantity called the truncated radius(trunc_rad)… this radius expands depending on the overlap. When I extract trunc_rad[i], I get the correct value but I get 0 for trunc_rad[j]. This messes up subsequent calculations that results in NaNs and eventually a segfault.
Here is the pertaining section of code where it fails where I calculate some quantities required to update the truncated radius:
double *trunc_rad = atom ->trunc_rad;
for (ii = 0; ii < inum; ii++) {
i = ilist[ii];
xtmp = x[i][0];
ytmp = x[i][1];
ztmp = x[i][2];
itype = type[i];
jlist = firstneigh[i];
cur_trunc_radi=trunc_rad[i];
radi = radius[i];
cur_radi=std::max(radi,cur_trunc_radi);
jnum = numneigh[i];
int count=0;
for (jj = 0; jj < jnum; jj++) {
j = jlist[jj];
j &= NEIGHMASK;
delx = xtmp - x[j][0];
dely = ytmp - x[j][1];
delz = ztmp - x[j][2];
xtmpj=x[j][0];
ytmpj=x[j][1];
ztmpj=x[j][2];
rsq = delx*delx + dely*dely + delz*delz;
cur_trunc_radj=trunc_rad[j];
radj = radius[j];
cur_radj=std::max(radj,cur_trunc_radj); //CODE FAILS HERE
radsum=cur_radi+cur_radj;
if (rsq < radsum*radsum) {
count+=1;
b=cur_radj+cur_radi;
b=sqrt(rsq);
s=(b+cur_radj+cur_radi)*0.5;
A=sqrt(s*(s-cur_radi)*(s-cur_radj)*(s-b));
h=2.0*A/b;
temp_h=(cur_radi*cur_radi-h*h)/(cur_radj*cur_radj-h*h);
if(b<radsum){
if(isnan(h)==1){
std::cout<<std::fixed<<std::setprecision(15)<<" "<<tag[i]<< " "<<tag[j]<<" "<<" xtmpi "<<xtmp<<" xympi "<<ytmp<<" ztmpi "<<ztmp<<" xtmpj "<<xtmpj<<" xympj "<<ytmpj<<" ztmpj "<<ztmpj<<"h in radloop is complex "<<h<<" b "<< b <<" s "<<s<< " A "<< A<< " s-cur_radi "<<s-cur_radi<<" s-cur_radj "<<s-cur_radj<< " s-b "<<s-b<<" curradi "<<cur_radi<< " curradj "<< cur_radj<< " radsum "<<radsum<< " rsq "<<rsq<< " radsum*radsum "<<radsum*radsum<<std::endl;
}
temp_r2=b/(1.0+sqrt(temp_h));
temp_r1=b-temp_r2;
temp_s1[i] += temp_r1;
temp_s2[i]+=temp_r1*temp_r1;
temp_n[i]+=1.0;
temp_s1[j] += temp_r2;
temp_s2[j]+=temp_r2*temp_r2;
temp_n[j]+=1.0;
}
}
}
}
My truncated radius in the input script is
9153: 0.004867644251833
9599: 0.004569340782181
However, during the loop in the above code, I get:
9153: 0.004867644251833
9599: 0
Not sure why this happens…
Any help is much appreciated! Can provide more code but I don’t know if that will be of help…
Best,